Annual Report 2023
Division of Cancer Immunology
Hiroyoshi Nishikawa, Shohei Koyama, Takuma Irie, Kota Itahashi, Kosuke Tanaka, Yasuko Tada, Shogo Kumagai, Genki Okumura, Sho Watanabe, Akihito Kawazoe, Junichiro Yuda, Megumi Fukuoka, Akito Fukuda, Yuki Koike, Kotaro Nomura, Yi-tzu Lin, Atsuo Sai, Sota Nakagawa, Rika Fujii, Daisuke Ito, Takuya Owari, Masaki Kondo, Tomohiro Iwasawa, Fumihiro Terasaki, Kazuhiro Kumagai, Tomoya Suzuki, Yasuki Adachi, Naoki Yuri, Takeo Naito, Sayuri Fukaya, Takehiko Manabe, Tamiyo Kobayashi, Sho Isoyama, Konomi Onagawa, Tomoka Takaku, Megumi Takemura, Megumi Hoshino, Chie Ozawa, Mizuho Kikuchi, Yoko Ohira, Sayuri Yoshimatsu
Introduction
Cancer immunotherapy, represented by immune checkpoint inhibitors (ICIs), has become one of the standard treatment options for various types of cancer. These ICIs are frequently administered in combination with conventional drugs in clinical settings. Various biomarkers such as PD-L1 expression in tumors and tumor mutation burden (TMB) have been identified to predict therapeutic responses, demonstrating their effectiveness in specific cancers. However, our understanding of markers based on stratification by immuno-genomic analysis of the tumor microenvironment (TME) that can accurately predict the therapeutic efficacy of ICIs remains limited. In addition, as the utilization of ICI therapy expands, the number of cases of long-term response and relapse after long-term treatment is increasing, and the mechanism(s) for long-term response and the resistance to ICIs needs to be clarified.
The Team and What We Do
To address these challenges, our laboratory collaborates with clinical teams to identify novel biomarkers related to clinical responses by ICIs, to elucidate the mechanisms involved in treatment resistance to ICIs, and to propose combination treatment strategies targeting these mechanisms. Focusing on regulatory T cells (Tregs), which play a major role in suppressing anti-tumor immunity, we analyze tumor tissues, blood, and stools before and after various treatments including ICIs, and comprehensively analyze both cancer cells and immune cells by multicolor flow cytometry, multiplex immunohistochemistry, CyTOF, single cell RNA/ATAC/TCR seq, whole exome analysis, and metagenomic analysis (Figure 1). In addition, we investigate the mechanisms of the phenomena observed in humans by transferring them to mouse models. We are also actively involved in collaborative research with pharmaceutical companies, aiming at introducing new biomarkers into clinical trials and developing new immunotherapies and combination cancer therapies.
Figure 1. We are investigating the dynamic immune state in cancer patients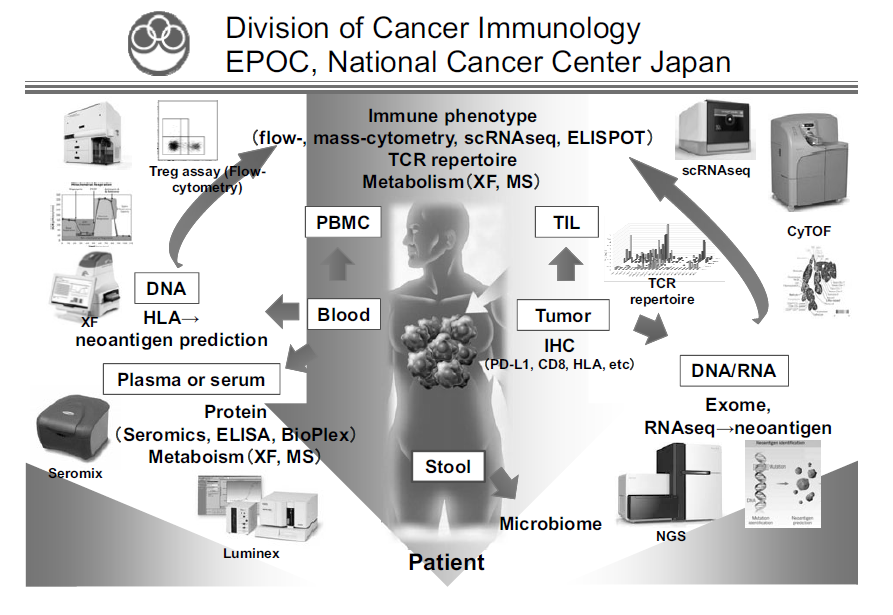

Research Activities
In collaboration with a clinical team led by the Department of Gastroenterology and Respiratory Medicine in the NCCHE and the NCCH, our laboratory conducted a research on biomarkers for predicting the therapeutic responses to anti-PD-1 antibodies, and identified that the balance of PD-1 expression between CD8+ T cells and Tregs is crucial to stratify responders from non-responders based on analysis of tumor-infiltrating T cells (Kumagai Shogo et al. Nat Immunol 2020) (Figure 2). Furthermore, we have analyzed the mechanisms of tumor immune evasion using human specimens and cancer-bearing mouse models and have reported on the mechanisms by which driver genes and cancer signals, including EGFR gene mutations, RHOA gene mutations, MYC gene amplification, and WNT/β-catenin signaling, edit the tumor micro-immune environment in an immunosuppressive manner via cytokines and metabolites (Sugiyama Eri et al. Sci Immunol. 2020, Kumagai Shogo et al. Immunity 2020, Takeuchi Yoshiko et al. Sci Immunol. 2021, Kumagai Shogo et al. Cancer Cell 2022). Including our research, the progress made in the field of tumor immunology over the past decade has shown that some driver genes and cancer signals not only induce cancer cell survival and proliferation, but also construct immunosuppressive networks at the same time, and we have proposed the concept of “immunogenomic cancer evolution” (Kumagai Shogo et al. Nat Rev Clin Oncol. 2024). Therefore, molecular targeted drugs that target these driver genes and cancer signals may not only directly kill cancer cells, but also block the immunosuppressive cancer network, and the importance of “immunogenomic precision medicine” is becoming clear.
Figure 2. Developing immuno-precision biomarkers and elucidating the mechanisms of PD-1 expression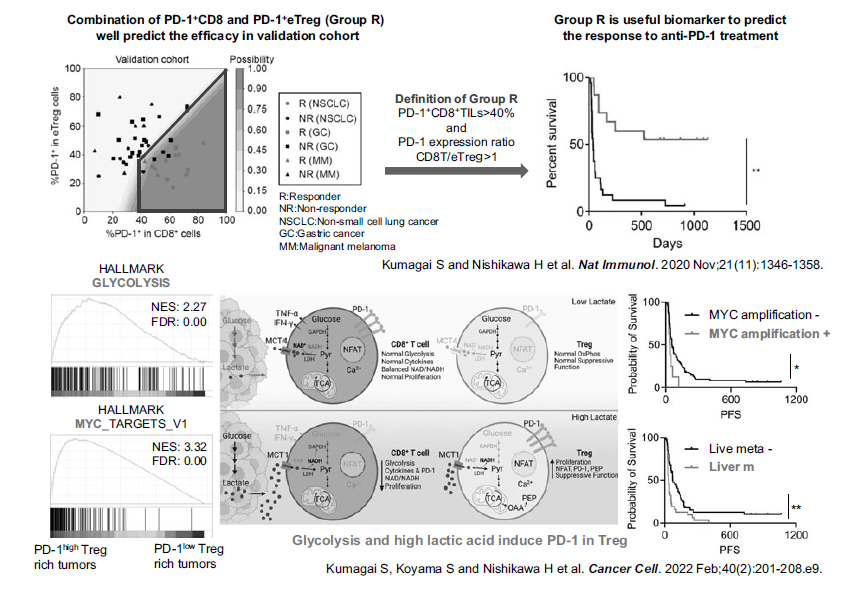

Clinical Trials
We have collaborated with several pharmaceutical companies in their clinical trials to elucidate biomarkers for predicting therapeutic responses and to evaluate immune status in the TME using our real-time immune monitoring.
Education
Many graduate school students are trained in our division, and young residents in the NCCHE are also trained to become physician scientists. After finishing their PhD course, some of the students continue their cancer immunology studies abroad.
Future Prospects
ICIs have undoubtedly opened a new era in cancer treatment. However, we still have several issues to be clarified, including acquired resistance and suppressive immune modulation by somatic mutations in cancer cells, to overcome the drawbacks of ICIs. In order to answer these questions, immunological analyses using clinical specimens are essential. Therefore, it is important to develop effective therapeutic strategies combining ICIs with other anti-cancer drugs and to identify biomarkers that can stratify responders from non-responders by clarifying the immune suppressive networks in the TME.
List of papers published in 2023
Journal
1. Takei S, Tanaka Y, Lin YT, Koyama S, Fukuoka S, Hara H, Nakamura Y, Kuboki Y, Kotani D, Kojima T, Bando H, Mishima S, Ueno T, Kojima S, Wakabayashi M, Sakamoto N, Kojima M, Kuwata T, Yoshino T, Nishikawa H, Mano H, Endo I, Shitara K, Kawazoe A. Multiomic molecular characterization of the response to combination immunotherapy in MSS/pMMR metastatic colorectal cancer. Journal for immunotherapy of cancer, 12:e008210, 2024
2. Shiraishi K, Takahashi A, Momozawa Y, Daigo Y, Kaneko S, Kawaguchi T, Kunitoh H, Matsumoto S, Horinouchi H, Goto A, Honda T, Shimizu K, Torasawa M, Takayanagi D, Saito M, Saito A, Ohe Y, Watanabe SI, Goto K, Tsuboi M, Tsuchihara K, Takata S, Aoi T, Takano A, Kobayashi M, Miyagi Y, Tanaka K, Suzuki H, Maeda D, Yamaura T, Matsuda M, Shimada Y, Mizuno T, Sakamoto H, Yoshida T, Goto Y, Yoshida T, Yamaji T, Sonobe M, Toyooka S, Yoneda K, Masago K, Tanaka F, Hara M, Fuse N, Nishizuka SS, Motoi N, Sawada N, Nishida Y, Kumada K, Takeuchi K, Tanno K, Yatabe Y, Sunami K, Hishida T, Miyazaki Y, Ito H, Amemiya M, Totsuka H, Nakayama H, Yokose T, Ishigaki K, Nagashima T, Ohtaki Y, Imai K, Takasawa K, Minamiya Y, Kobayashi K, Okubo K, Wakai K, Shimizu A, Yamamoto M, Iwasaki M, Matsuda K, Inazawa J, Shiraishi Y, Nishikawa H, Murakami Y, Kubo M, Matsuda F, Kamatani Y, Hamamoto R, Matsuo K, Kohno T. Identification of telomere maintenance gene variations related to lung adenocarcinoma risk by genome-wide association and whole genome sequencing analyses. Cancer communications (London, England), 44:287-293, 2024
3. Hirai T, Naito Y, Koyama S, Nakanishi Y, Masuhiro K, Izumi M, Kuge T, Naito M, Mizuno Y, Yamaguchi Y, Kang S, Yaga M, Futami Y, Nojima S, Nishide M, Morita T, Kato Y, Tsuda T, Takemoto N, Kinugasa-Katayama Y, Aoshi T, Villa JK, Yamashita K, Enokida T, Hoshi Y, Matsuura K, Tahara M, Takamatsu H, Takeda Y, Inohara H, Kumanogoh A. Sema6D forward signaling impairs T cell activation and proliferation in head and neck cancer. JCI insight, 9:e166349, 2024
4. Kondo M, Kumagai S, Nishikawa H. Metabolic advantages of regulatory T cells dictated by cancer cells. International immunology, 36:75-86, 2024
5. Nishikawa H. Establishment of immune suppression by cancer cells in the tumor microenvironment. Proceedings of the Japan Academy. Series B, Physical and biological sciences, 100:114-122, 2024
6. Zhou W, Kawashima S, Ishino T, Kawase K, Ueda Y, Yamashita K, Watanabe T, Kawazu M, Dansako H, Suzuki Y, Nishikawa H, Inozume T, Nagasaki J, Togashi Y. Stem-like progenitor and terminally differentiated T(FH)-like CD4(+) T cell exhaustion in the tumor microenvironment. Cell reports, 43:113797, 2024
7. Kumagai S, Itahashi K, Nishikawa H. Regulatory T cell-mediated immunosuppression orchestrated by cancer: towards an immuno-genomic paradigm for precision medicine. Nature reviews. Clinical oncology, 21:337-353, 2024
8. Terasaki F, Sugiura T, Okamura Y, Ashida R, Ohgi K, Yamada M, Ohtsuka S, Uesaka K. Benefit of lymph node dissection for perihilar and distal cholangiocarcinoma according to lymph node stations. Journal of hepato-biliary-pancreatic sciences, 31:251-261, 2024
9. Okuma HS, Watanabe K, Tsuchihashi K, Machida R, Sadachi R, Hirakawa A, Ariyama H, Kanai M, Kamikura M, Anjo K, Hiramitsu A, Sekine S, Okita N, Mano H, Nishikawa H, Nakamura K, Yonemori K. Phase II Trial of Nivolumab in Metastatic Rare Cancer with dMMR or MSI-H and Relation with Immune Phenotypic Analysis (the ROCK Trial). Clinical cancer research, 29:5079-5086, 2023
10. Fukuda A, Yoshida T. Treatment of advanced ALK-rearranged NSCLC following second-generation ALK-TKI failure. Expert review of anticancer therapy, 23:1157-1167, 2023
11. Watanabe S, Shimoi T, Nishikawa T, Kawachi A, Okuma HS, Tokura M, Yazaki S, Mizoguchi C, Arakaki M, Saito A, Kita S, Yamamoto K, Kojima Y, Sudo K, Noguchi E, Yoshida A, Kawai A, Fujiwara Y, Yonemori K. Lymphocyte-to-monocyte ratio as a prognostic and potential tumor microenvironment indicator in advanced soft tissue sarcoma treated with first-line doxorubicin therapy. Scientific reports, 13:10734, 2023
12. Habu T, Kumanishi R, Ogata T, Fujisawa T, Mishima S, Kotani D, Kadowaki S, Nakamura M, Hojo H, Fujiwara H, Kumagai S, Koyama S, Fujita T, Kinoshita T, Nishikawa H, Yano T, Tajika M, Muro K, Mitsunaga S, Kojima T, Bando H. Complete response to definitive chemoradiotherapy in unresectable locally advanced esophageal squamous cell carcinoma. Esophagus, 20:533-540, 2023
13. Kumagai S, Koyama S, Shitara K. Precise patient stratification in esophageal cancer: Biomarkers for immunochemotherapy. Cancer cell, 41:1199-1201, 2023
14. Du J, Kageyama SI, Yamashita R, Tanaka K, Okumura M, Motegi A, Hojo H, Nakamura M, Hirata H, Sunakawa H, Kotani D, Yano T, Kojima T, Hamaya Y, Kojima M, Nakamura Y, Suzuki A, Suzuki Y, Tsuchihara K, Akimoto T. Transposable elements potentiate radiotherapy-induced cellular immune reactions via RIG-I-mediated virus-sensing pathways. Communications biology, 6:818, 2023
15. Morita TY, Yu J, Kashima Y, Kamata R, Yamamoto G, Minamide T, Mashima C, Yoshiya M, Sakae Y, Yamauchi T, Hakozaki Y, Kageyama SI, Nakamura A, Lightcap E, Tanaka K, Niu H, Kannan K, Ohashi A. CDC7 inhibition induces replication stress-mediated aneuploid cells with an inflammatory phenotype sensitizing tumors to immune checkpoint blockade. Nature communications, 14:7490, 2023
16. Hiramatsu H, Nosaka K, Kusumoto S, Nakano N, Choi I, Yoshimitsu M, Imaizumi Y, Hidaka M, Sasaki H, Makiyama J, Ohtsuka E, Jo T, Ogata M, Ito A, Yonekura K, Tatetsu H, Kato T, Kawakita T, Suehiro Y, Ishitsuka K, Iida S, Matsutani T, Nishikawa H, Utsunomiya A, Ueda R, Ishida T. Landscape of immunoglobulin heavy chain γ gene class switch recombination in patients with adult T-cell leukemia-lymphoma. Haematologica, 108:1173-1178, 2023
17. Kato S, Maeda Y, Sugiyama D, Watanabe K, Nishikawa H, Hinohara K. The cancer epigenome: Non-cell autonomous player in tumor immunity. Cancer science, 114:730-740, 2023
18. Ito M, Iwama S, Sugiyama D, Yasuda Y, Okuji T, Kobayashi T, Zhou X, Yamagami A, Onoue T, Miyata T, Sugiyama M, Hagiwara D, Suga H, Banno R, Nishikawa H, Arima H. Anti-tumor effects of anti-programmed cell death-1 antibody treatment are attenuated in streptozotocin-induced diabetic mice. Scientific reports, 13:5939, 2023
19. Harusato A, Seo W, Abo H, Nakanishi Y, Nishikawa H, Itoh Y. Impact of particulate microplastics generated from polyethylene terephthalate on gut pathology and immune microenvironments. iScience, 26:106474, 2023
20. Nishiwaki S, Sugiura I, Sato T, Kobayashi M, Osaki M, Sawa M, Adachi Y, Okabe M, Saito S, Morishita T, Kohno A, Nishiyama T, Iida H, Kurahashi S, Kuwatsuka Y, Sugiyama D, Ito S, Nishikawa H, Kiyoi H. Autologous peripheral blood stem cell transplantation for Philadelphia chromosome-positive acute lymphoblastic leukemia is safe but poses challenges for long-term maintenance of molecular remission: Results of the Auto-Ph17 study. EJHaem, 4:358-369, 2023
21. Barakat C, Inagaki Y, Mizuno S, Nishio N, Katsuyama N, Sato Y, Kobayashi M, Ozeki K, Iida H, Tomita A, Sawa M, Demachi-Okamura A, Takahashi Y, Nishikawa H, Akatsuka Y. Development of TCR-T cell therapy targeting mismatched HLA-DPB1 for relapsed leukemia after allogeneic transplantation. International journal of hematology, 118:252-266, 2023
22. Hiramatsu H, Yokomori R, Shengyi L, Tanaka N, Mori S, Kiyotani K, Gotoh O, Kusumoto S, Nakano N, Suehiro Y, Ito A, Choi I, Ohtsuka E, Hidaka M, Nosaka K, Yoshimitsu M, Imaizumi Y, Iida S, Utsunomiya A, Noda T, Nishikawa H, Ueda R, Sanda T, Ishida T. Clinical landscape of TP73 structural variants in ATL patients. Leukemia, 37:2502-2506, 2023
23. Harusato A, Seo W, Abo H, Nakanishi Y, Nishikawa H, Itoh Y. Protocol for acquiring samples to assess the impact of microplastics on immune microenvironments in the mouse intestine. STAR protocols, 4:102648, 2023
24. Xiao M, Kondo S, Nomura M, Kato S, Nishimura K, Zang W, Zhang Y, Akashi T, Viny A, Shigehiro T, Ikawa T, Yamazaki H, Fukumoto M, Tanaka A, Hayashi Y, Koike Y, Aoyama Y, Ito H, Nishikawa H, Kitamura T, Kanai A, Yokoyama A, Fujiwara T, Goyama S, Noguchi H, Lee SC, Toyoda A, Hinohara K, Abdel-Wahab O, Inoue D. BRD9 determines the cell fate of hematopoietic stem cells by regulating chromatin state. Nature communications, 14:8372, 2023
25. Katsuyama N, Kawase T, Barakat C, Mizuno S, Tomita A, Ozeki K, Nishio N, Sato Y, Kajiya R, Shiraishi K, Takahashi Y, Ichinohe T, Nishikawa H, Akatsuka Y. T cell receptor-engineered T cells derived from target human leukocyte antigen-DPB1-specific T cell can be a potential tool for therapy against leukemia relapse following allogeneic hematopoietic cell transplantation. Nagoya journal of medical science, 85:779-796, 2023
26. Calthorpe L, Rashidian N, Cacciaguerra AB, Conroy PC, Hibi T, Hilal MA, Hoffman D, Park KM, Wang J, Adam MA, Alseidi A. Using the Comprehensive Complication Index to Rethink the ISGLS Criteria for Post-hepatectomy Liver Failure in an International Cohort of Major Hepatectomies. Annals of surgery, 277:e592-e596, 2023
27. Naito Y, Koyama S, Masuhiro K, Hirai T, Uenami T, Inoue T, Osa A, Machiyama H, Watanabe G, Sax N, Villa J, Kinugasa-Katayama Y, Nojima S, Yaga M, Hosono Y, Okuzaki D, Satoh S, Tsuda T, Nakanishi Y, Suga Y, Morita T, Fukushima K, Nishide M, Shiroyama T, Miyake K, Iwahori K, Hirata H, Nagatomo I, Yano Y, Tamiya M, Kumagai T, Takemoto N, Inohara H, Yamasaki S, Yamashita K, Aoshi T, Akbay EA, Hosen N, Shintani Y, Takamatsu H, Mori M, Takeda Y, Kumanogoh A. Tumor-derived semaphorin 4A improves PD-1-blocking antibody efficacy by enhancing CD8(+) T cell cytotoxicity and proliferation. Science advances, 9:eade0718, 2023
28. Wang JJ, Feng J, Gomes C, Calthorpe L, Ashraf Ganjouei A, Romero-Hernandez F, Benedetti Cacciaguerra A, Hibi T, Adam MA, Alseidi A, Abu Hilal M, Rashidian N. Development and Validation of Prediction Models and Risk Calculators for Posthepatectomy Liver Failure and Postoperative Complications Using a Diverse International Cohort of Major Hepatectomies. Annals of surgery, 278:976-984, 2023
29. Terasaki F, Hirakawa S, Tachimori H, Sugiura T, Nanashima A, Komatsu S, Miyata H, Kakeji Y, Kitagawa Y, Nakamura M, Endo I. Morbidity after left trisectionectomy for hepato-biliary malignancies: An analysis of the National Clinical Database of Japan. Journal of hepato-biliary-pancreatic sciences, 30:1304-1315, 2023
30. Manabe T, Mori M, Kanayama M, Kuwata T, Takenaka M, Kuroda K, Fukumitsu S, Murakami Y, Aoki T, Nakayama T, Tanaka F. Successful resection of a hypervascular paravertebral solitary fibrous tumor of pleura preserving the artery of Adamkiewicz, which was detected on preoperative angiography. General Thoracic and Cardiovascular Surgery Cases, 2:2023
