Annual Report 2023
Division of Cancer Immunology
Hiroyoshi Nishikawa, Yuka Maeda, Kosuke Tanaka, Keisuke Watanabe, Hitomi Nishinakamura, Kota Itahashi, Priya Saju, Sana Hibino, Akihito Nagata, Nanae Shimoyama, Yuki Ishige, Kyo Ishin, May Thinzar Hlaing, Abdullah AimiNaim
Introduction
Cancer immunotherapies such as immune checkpoint inhibitors (ICIs) or chimeric antigen receptor T (CAR-T) cell or T- cell receptor T (TCR-T) cell therapies have become an established therapy for some cancers. However, not a few patients experience relapse or resistant disease. Cancers often create a complex immune suppressive environment that can impair the immune reaction, thereby impairing the efficacy of ICIs or adoptive transferred gene-modified T cells. Therefore, there is an urgent demand for elucidating the mechanisms of resistance and identifying reliable biomarkers to predict their efficacy to make immunotherapies much more effective in clinical settings.
The Team and What We Do
We have been collecting clinical samples from cancer patients treated with ICI or gene-modified cell therapies and intensively investigating the mechanisms of cancer immunoreactions using multi-omics analysis including multicolor flow cytometry, mass cytometry, imaging mass cytometry, single-cell transcriptome analysis, proteomics, and metabolomics (Figure 1). We are also developing first-in-class gene-modified, cell-based immunotherapies using CAR-T cell or TCR-T cell platforms targeting the above mechanisms of resistance (Figure 2).
Figure 1. Integrated multi-omics analyses on patient-derived tumor/immune cell samples.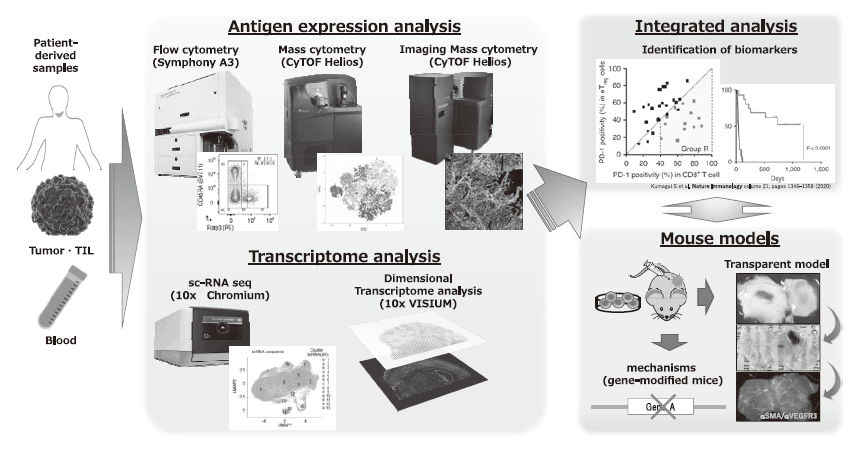

Figure 2. Drug discovery using chimeric antigen receptor T cell (CAR-T cell) format.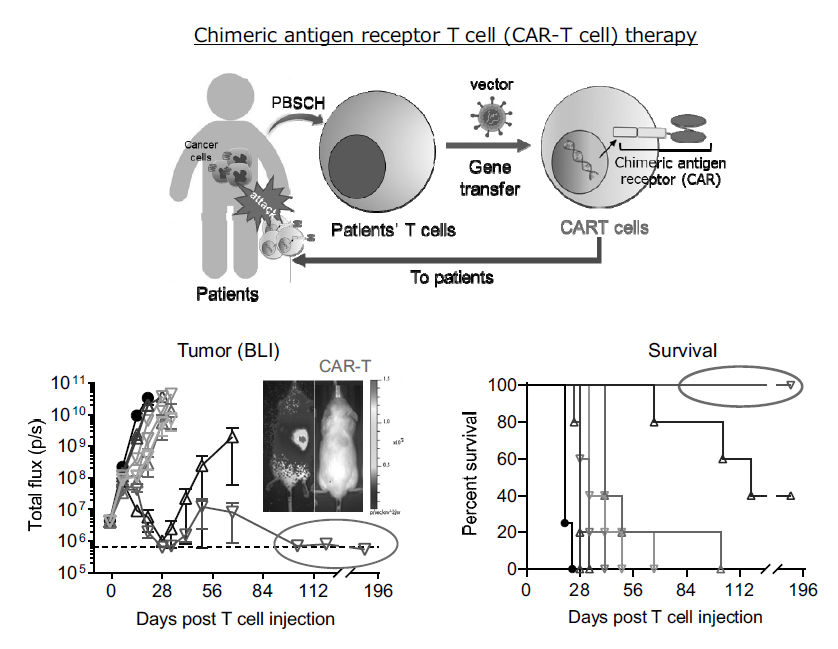

Research Activities
We have elucidated critical mechanisms that impair the efficacy of cancer immunotherapies using patient-derived samples pre- and post-treatment with treatment of ICIs or gene-modified cell therapies. In addition, we have elucidated the differentiation process and epigenetic profiles of regulatory T cells (Treg cells) in the tumor microenvironment (TME). BATF was identified as a key regulator, which leveraged Treg cell differentiation through epigenetically controlling activation-associated gene expression, resulting in the robustness of Treg cells in the TME. We are also developing novel CAR-T cells that overcome multiple immunosuppressive factors to improve the treatment outcome of adoptive cell therapies in solid tumors. Some of them are under pre-clinical testing aiming for first-in-human trials. In addition, we have developed a combination therapy with a drug and anti-PD-1 Ab. This drug activates innate immunity via Toll-like receptor signaling. For solid tumors that do not respond to anti-PD-1 antibody therapy, we analyzed the anti-tumor immune response by stimulating the innate immune response in tumor-bearing mice model. The phenotype and proportion of the various immune cells that infiltrate tumors have been identified, which could lead to the development of a novel combination cancer immunotherapy.
Clinical Trials
An investigator-initiated first-in-human trial of our CAR-T cell therapy for relapsed or refractory T cell malignancies is under preparation.
Education
We are hosting graduate students from several universities with partnership agreements, and young residents from the National Cancer Center Hospital for training in translational research. We strongly support their career development, and many alumni are leading the immunology field in a variety of posts in the clinic, academia, or paratheatrical companies.
Future Prospects
We will further investigate the mechanisms involved in tumor immune suppression or tumor immune evasion in various cancers with a broad view into such areas as immunology and genetics, using new approaches including special and dynamic analysis of the transcriptome or immune cell phenotypes, and transparent mouse tissue models. Our findings will lead to drug discoveries to overcome resistance and to identify of biomarkers in immune therapies. We are planning a first-in-human trial of CAR-T cell therapy for T cell malignancies that has been developed in our lab.
List of papers published in 2023
Journal
1. Takei S, Tanaka Y, Lin YT, Koyama S, Fukuoka S, Hara H, Nakamura Y, Kuboki Y, Kotani D, Kojima T, Bando H, Mishima S, Ueno T, Kojima S, Wakabayashi M, Sakamoto N, Kojima M, Kuwata T, Yoshino T, Nishikawa H, Mano H, Endo I, Shitara K, Kawazoe A. Multiomic molecular characterization of the response to combination immunotherapy in MSS/pMMR metastatic colorectal cancer. Journal for immunotherapy of cancer, 12:e008210, 2024
2. Shiraishi K, Takahashi A, Momozawa Y, Daigo Y, Kaneko S, Kawaguchi T, Kunitoh H, Matsumoto S, Horinouchi H, Goto A, Honda T, Shimizu K, Torasawa M, Takayanagi D, Saito M, Saito A, Ohe Y, Watanabe SI, Goto K, Tsuboi M, Tsuchihara K, Takata S, Aoi T, Takano A, Kobayashi M, Miyagi Y, Tanaka K, Suzuki H, Maeda D, Yamaura T, Matsuda M, Shimada Y, Mizuno T, Sakamoto H, Yoshida T, Goto Y, Yoshida T, Yamaji T, Sonobe M, Toyooka S, Yoneda K, Masago K, Tanaka F, Hara M, Fuse N, Nishizuka SS, Motoi N, Sawada N, Nishida Y, Kumada K, Takeuchi K, Tanno K, Yatabe Y, Sunami K, Hishida T, Miyazaki Y, Ito H, Amemiya M, Totsuka H, Nakayama H, Yokose T, Ishigaki K, Nagashima T, Ohtaki Y, Imai K, Takasawa K, Minamiya Y, Kobayashi K, Okubo K, Wakai K, Shimizu A, Yamamoto M, Iwasaki M, Matsuda K, Inazawa J, Shiraishi Y, Nishikawa H, Murakami Y, Kubo M, Matsuda F, Kamatani Y, Hamamoto R, Matsuo K, Kohno T. Identification of telomere maintenance gene variations related to lung adenocarcinoma risk by genome-wide association and whole genome sequencing analyses. Cancer communications (London, England), 44:287-293, 2024
3. Kondo M, Kumagai S, Nishikawa H. Metabolic advantages of regulatory T cells dictated by cancer cells. International immunology, 36:75-86, 2024
4. Nishikawa H. Establishment of immune suppression by cancer cells in the tumor microenvironment. Proceedings of the Japan Academy. Series B, Physical and biological sciences, 100:114-122, 2024
5. Zhou W, Kawashima S, Ishino T, Kawase K, Ueda Y, Yamashita K, Watanabe T, Kawazu M, Dansako H, Suzuki Y, Nishikawa H, Inozume T, Nagasaki J, Togashi Y. Stem-like progenitor and terminally differentiated T(FH)-like CD4(+) T cell exhaustion in the tumor microenvironment. Cell reports, 43:113797, 2024
6. Kumagai S, Itahashi K, Nishikawa H. Regulatory T cell-mediated immunosuppression orchestrated by cancer: towards an immuno-genomic paradigm for precision medicine. Nature reviews. Clinical oncology, 21:337-353, 2024
7. Okuma HS, Watanabe K, Tsuchihashi K, Machida R, Sadachi R, Hirakawa A, Ariyama H, Kanai M, Kamikura M, Anjo K, Hiramitsu A, Sekine S, Okita N, Mano H, Nishikawa H, Nakamura K, Yonemori K. Phase II Trial of Nivolumab in Metastatic Rare Cancer with dMMR or MSI-H and Relation with Immune Phenotypic Analysis (the ROCK Trial). Clinical cancer research, 29:5079-5086, 2023
8. Habu T, Kumanishi R, Ogata T, Fujisawa T, Mishima S, Kotani D, Kadowaki S, Nakamura M, Hojo H, Fujiwara H, Kumagai S, Koyama S, Fujita T, Kinoshita T, Nishikawa H, Yano T, Tajika M, Muro K, Mitsunaga S, Kojima T, Bando H. Complete response to definitive chemoradiotherapy in unresectable locally advanced esophageal squamous cell carcinoma. Esophagus, 20:533-540, 2023
9. Du J, Kageyama SI, Yamashita R, Tanaka K, Okumura M, Motegi A, Hojo H, Nakamura M, Hirata H, Sunakawa H, Kotani D, Yano T, Kojima T, Hamaya Y, Kojima M, Nakamura Y, Suzuki A, Suzuki Y, Tsuchihara K, Akimoto T. Transposable elements potentiate radiotherapy-induced cellular immune reactions via RIG-I-mediated virus-sensing pathways. Communications biology, 6:818, 2023
10. Morita TY, Yu J, Kashima Y, Kamata R, Yamamoto G, Minamide T, Mashima C, Yoshiya M, Sakae Y, Yamauchi T, Hakozaki Y, Kageyama SI, Nakamura A, Lightcap E, Tanaka K, Niu H, Kannan K, Ohashi A. CDC7 inhibition induces replication stress-mediated aneuploid cells with an inflammatory phenotype sensitizing tumors to immune checkpoint blockade. Nature communications, 14:7490, 2023
11. Hiramatsu H, Nosaka K, Kusumoto S, Nakano N, Choi I, Yoshimitsu M, Imaizumi Y, Hidaka M, Sasaki H, Makiyama J, Ohtsuka E, Jo T, Ogata M, Ito A, Yonekura K, Tatetsu H, Kato T, Kawakita T, Suehiro Y, Ishitsuka K, Iida S, Matsutani T, Nishikawa H, Utsunomiya A, Ueda R, Ishida T. Landscape of immunoglobulin heavy chain γ gene class switch recombination in patients with adult T-cell leukemia-lymphoma. Haematologica, 108:1173-1178, 2023
12. Kato S, Maeda Y, Sugiyama D, Watanabe K, Nishikawa H, Hinohara K. The cancer epigenome: Non-cell autonomous player in tumor immunity. Cancer science, 114:730-740, 2023
13. Ito M, Iwama S, Sugiyama D, Yasuda Y, Okuji T, Kobayashi T, Zhou X, Yamagami A, Onoue T, Miyata T, Sugiyama M, Hagiwara D, Suga H, Banno R, Nishikawa H, Arima H. Anti-tumor effects of anti-programmed cell death-1 antibody treatment are attenuated in streptozotocin-induced diabetic mice. Scientific reports, 13:5939, 2023
14. Watanabe K, Gomez AM, Kuramitsu S, Siurala M, Da T, Agarwal S, Song D, Scholler J, Rotolo A, Posey AD, Rook AH, Haun PL, Ruella M, Young RM, June CH. Identifying highly active anti-CCR4 CAR T cells for the treatment of T-cell lymphoma. Blood advances, 7:3416-3430, 2023
15. Harusato A, Seo W, Abo H, Nakanishi Y, Nishikawa H, Itoh Y. Impact of particulate microplastics generated from polyethylene terephthalate on gut pathology and immune microenvironments. iScience, 26:106474, 2023
16. Nishiwaki S, Sugiura I, Sato T, Kobayashi M, Osaki M, Sawa M, Adachi Y, Okabe M, Saito S, Morishita T, Kohno A, Nishiyama T, Iida H, Kurahashi S, Kuwatsuka Y, Sugiyama D, Ito S, Nishikawa H, Kiyoi H. Autologous peripheral blood stem cell transplantation for Philadelphia chromosome-positive acute lymphoblastic leukemia is safe but poses challenges for long-term maintenance of molecular remission: Results of the Auto-Ph17 study. EJHaem, 4:358-369, 2023
17. Hibino S, Eto S, Hangai S, Endo K, Ashitani S, Sugaya M, Osawa T, Soga T, Taniguchi T, Yanai H. Tumor cell-derived spermidine is an oncometabolite that suppresses TCR clustering for intratumoral CD8(+) T cell activation. Proceedings of the National Academy of Sciences of the United States of America, 120:e2305245120, 2023
18. Barakat C, Inagaki Y, Mizuno S, Nishio N, Katsuyama N, Sato Y, Kobayashi M, Ozeki K, Iida H, Tomita A, Sawa M, Demachi-Okamura A, Takahashi Y, Nishikawa H, Akatsuka Y. Development of TCR-T cell therapy targeting mismatched HLA-DPB1 for relapsed leukemia after allogeneic transplantation. International journal of hematology, 118:252-266, 2023
19. Kato M, Maeda K, Nakahara R, Hirose H, Kondo A, Aki S, Sugaya M, Hibino S, Nishida M, Hasegawa M, Morita H, Ando R, Tsuchida R, Yoshida M, Kodama T, Yanai H, Shimamura T, Osawa T. Acidic extracellular pH drives accumulation of N1-acetylspermidine and recruitment of protumor neutrophils. PNAS nexus, 2:pgad306, 2023
20. Hiramatsu H, Yokomori R, Shengyi L, Tanaka N, Mori S, Kiyotani K, Gotoh O, Kusumoto S, Nakano N, Suehiro Y, Ito A, Choi I, Ohtsuka E, Hidaka M, Nosaka K, Yoshimitsu M, Imaizumi Y, Iida S, Utsunomiya A, Noda T, Nishikawa H, Ueda R, Sanda T, Ishida T. Clinical landscape of TP73 structural variants in ATL patients. Leukemia, 37:2502-2506, 2023
21. Harusato A, Seo W, Abo H, Nakanishi Y, Nishikawa H, Itoh Y. Protocol for acquiring samples to assess the impact of microplastics on immune microenvironments in the mouse intestine. STAR protocols, 4:102648, 2023
22. Xiao M, Kondo S, Nomura M, Kato S, Nishimura K, Zang W, Zhang Y, Akashi T, Viny A, Shigehiro T, Ikawa T, Yamazaki H, Fukumoto M, Tanaka A, Hayashi Y, Koike Y, Aoyama Y, Ito H, Nishikawa H, Kitamura T, Kanai A, Yokoyama A, Fujiwara T, Goyama S, Noguchi H, Lee SC, Toyoda A, Hinohara K, Abdel-Wahab O, Inoue D. BRD9 determines the cell fate of hematopoietic stem cells by regulating chromatin state. Nature communications, 14:8372, 2023
23. Katsuyama N, Kawase T, Barakat C, Mizuno S, Tomita A, Ozeki K, Nishio N, Sato Y, Kajiya R, Shiraishi K, Takahashi Y, Ichinohe T, Nishikawa H, Akatsuka Y. T cell receptor-engineered T cells derived from target human leukocyte antigen-DPB1-specific T cell can be a potential tool for therapy against leukemia relapse following allogeneic hematopoietic cell transplantation. Nagoya journal of medical science, 85:779-796, 2023
