Annual Report 2024
Division of Translational Informatics
Katsuya Tsuchihara, Riu Yamashita, Hiroko Nagayama, Shunsuke Sakai, Ryosuke Nomura, Masaki Ohira, Kyoko Kurihara
Introduction
We performed multi-omics data analysis on large-scale datasets to advance translational informatics.
The Team and What We Do
We participated in the MONSTAR-SCREEN1–3 projects, where we collected, organized, and analyzed the data. In addition, using data from the GALAXY trial in combination with the JoGo survey sequence database, we analyzed germline mutations associated with disease-free survival in colorectal cancer and investigated their biological implications.
We have also been engaged in spatial omics analysis, developing tools and performing data interpretation. In particular, we focused on creating methods for location-aware analysis, an area where current spatial omics approaches remain limited.
Furthermore, using a dataset of healthy individuals provided by the Tohoku Medical Megabank Organization (ToMMo), we constructed a database for Clonal Hematopoiesis of Indeterminate Potential (CHIP) and performed comparative analyses with cancer patients.
In addition, we conducted microbiome research, conducting association analyses between gut and oral microbiota and various cancers, including gynecological cancers, breast cancer, glioma, and head and neck cancer.
Research Activities
In ovarian cancer patients, specific gut microbiota were demonstrated to correlate with the therapeutic efficacy of PARP inhibitors. Furthermore, the spatial omics data analysis method SpatialKNifeY (SKNY) was developed, enabling the identification of microenvironment characteristics including the tumor surrounding environment (Figure 1).
Figure 1. Results of analysis performed with SpatialKnifeY (SKNY) 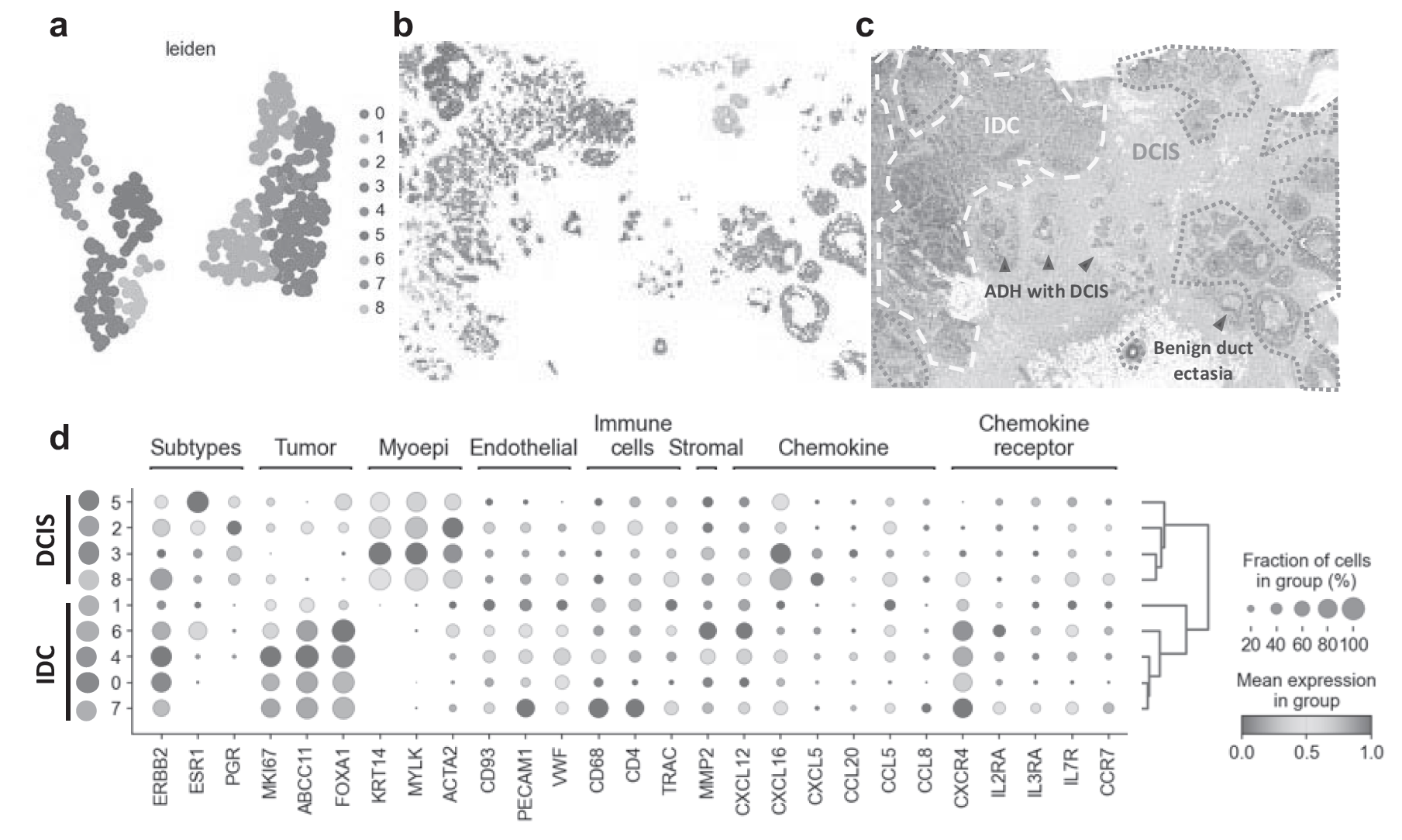

As an application, oxygen saturation imaging was shown to visualize and analyze tumor heterogeneity in gastric cancer. Furthermore, mathematical modeling was used to predict the optimal combination and timing of immune checkpoint inhibitors with radiation therapy. In collaboration with ToMMo, a series of pipelines was reported that utilize MassARRAY and genomic information to detect and correct sample misidentification in biobanks.
Education
Approximately 20 bioinformatics study sessions were held annually, with about 15 participants in total. In addition, lectures and hands-on training sessions were conducted at Tokyo University of Science and the University of Tokyo. Efforts were also made to promote AI within the Center by supporting participants in AI workshops and contests. Furthermore, three-month domestic study programs were organized for students from Osaka University, Gifu University, and Kumamoto University, providing research guidance through bioinformatics.
Future Prospects
We will continue to develop tools and extract biological and medical insights from large-scale, multi-layered omics datasets obtained from projects such as MONSTAR.
List of papers published in 2024
Journal
1. Nakamura Y, Ozaki H, Ueno M, Komatsu Y, Yuki S, Esaki T, Taniguchi H, Sunakawa Y, Yamaguchi K, Kato K, Denda T, Nishina T, Takahashi N, Satoh T, Yasui H, Satake H, Oki E, Kato T, Ohta T, Matsuhashi N, Goto M, Okano N, Ohtsubo K, Yamazaki K, Yamashita R, Iida N, Yuasa M, Bando H, Yoshino T. Targeted therapy guided by circulating tumor DNA analysis in advanced gastrointestinal tumors. Nature medicine, 31:165-175, 2025
2. Umemura S, Udagawa H, Ikeda T, Murakami H, Daga H, Toyozawa R, Kozuki T, Sakakibara-Konishi J, Ohe Y, Morise M, Kato T, Shingyoji M, Hara S, Furuya N, Teranishi S, Takata S, Miyamoto S, Nakachi I, Wakabayashi M, Nomura S, Sato A, Ishii G, Tsuchihara K, Sugiyama E, Kirita K, Sakai T, Shibata Y, Izumi H, Nosaki K, Zenke Y, Matsumoto S, Yoh K, Niho S, Goto K. Clinical Significance of a Prospective Large Genomic Screening for SCLC: The Genetic Classification and a Biomarker-Driven Phase 2 Trial of Gedatolisib. Journal of thoracic oncology, 20:177-193, 2025
3. Okazawa-Sakai M, Sakai SA, Hyodo I, Horasawa S, Sawada K, Fujisawa T, Yamamoto Y, Boku S, Hayasaki Y, Isobe M, Shintani D, Hasegawa K, Egawa-Takata T, Ito K, Ihira K, Watari H, Takehara K, Yagi H, Kato K, Chiyoda T, Harano K, Nakamura Y, Yamashita R, Yoshino T, Aoki D. Gut microbiome associated with PARP inhibitor efficacy in patients with ovarian cancer. Journal of gynecologic oncology, 36:e38, 2025
4. Sakai SA, Saeki K, Chi S, Hamaya Y, Du J, Nakamura M, Hojo H, Kojima T, Nakamura Y, Bando H, Kojima M, Suzuki A, Suzuki Y, Akimoto T, Tsuchihara K, Haeno H, Yamashita R, Kageyama SI. Mathematical Modeling Predicts Optimal Immune Checkpoint Inhibitor and Radiotherapy Combinations and Timing of Administration. Cancer immunology research, 13:353-364, 2025
5. Imai M, Nakamura Y, Shin S, Okamoto W, Kato T, Esaki T, Kato K, Komatsu Y, Yuki S, Masuishi T, Nishina T, Sawada K, Sato A, Kuwata T, Yamashita R, Fujisawa T, Bando H, Ock CY, Fujii S, Yoshino T. Artificial Intelligence-Powered Human Epidermal Growth Factor Receptor 2 and Tumor Microenvironment Analysis in Human Epidermal Growth Factor Receptor 2-Amplified Metastatic Colorectal Cancer: Exploratory Analysis of Phase II TRIUMPH Trial. JCO precision oncology, 9:e2400385, 2025
6. Sakai SA, Nomura R, Nagasawa S, Chi S, Suzuki A, Suzuki Y, Imai M, Nakamura Y, Yoshino T, Ishikawa S, Tsuchihara K, Kageyama SI, Yamashita R. SpatialKNifeY (SKNY): Extending from spatial domain to surrounding area to identify microenvironment features with single-cell spatial omics data. PLoS computational biology, 21:e1012854, 2025
7. Ishihara H, Yamashita R, Ishiyama R, Ikeda T, Fukuda H, Yoshida K, Hirai T, Iizuka J, Kondo T, Nagashima Y, Takagi T. Genome-wide transcriptome and DNA methylome profiling of acquired cystic disease-associated renal cell carcinoma. Pathology, 57:495-501, 2025
8. Minamide T, Minakata N, Yamashita R, Sakashita S, Yoda Y, Ohashi A, Aoshima M, Kobayashi S, Yano T. Oxygen saturation imaging elucidates tumor heterogeneity in gastric cancer. DEN open, 5:e70077, 2025
9. Kudo H, Ishida N, Nobukuni T, Aoki Y, Saito S, Nishijima I, Terakawa T, Yamamoto M, Minegishi N, Yamashita R, Kumada K. Detection and Correction of Sample Misidentifications in a Biobank Using the MassARRAY System and Genomic Information. Biopreservation and biobanking, 22:373-382, 2024
10. Hashimoto T, Nakamura Y, Fujisawa T, Imai M, Shibuki T, Iida N, Ozaki H, Nonomura N, Morizane C, Iwata H, Okano S, Yamagami W, Yamazaki N, Kadowaki S, Taniguchi H, Ueno M, Boku S, Oki E, Komatsu Y, Yuki S, Makiyama A, Otsuka T, Hara H, Okano N, Nishina T, Sakamoto Y, Miki I, Kobayashi S, Yuda J, Kageyama SI, Nagamine M, Sakashita S, Sakamoto N, Yamashita R, Koga Y, Bando H, Ishii G, Kuwata T, Park WY, Ohtsu A, Yoshino T. The SCRUM-MONSTAR Cancer-Omics Ecosystem: Striving for a Quantum Leap in Precision Medicine. Cancer discovery, 14:2243-2261, 2024
11. Watanabe M, Haeno H, Mimaki S, Tsuchihara K. Multistage carcinogenesis in occupational cholangiocarcinoma: the impact of clonal expansion and risk estimation. Genes and environment, 46:21, 2024
12. Hasegawa S, Shoji Y, Kato M, Elzawahry A, Nagai M, Gi M, Suzuki S, Wanibuchi H, Mimaki S, Tsuchihara K, Totsuka Y. Whole Genome Sequencing Analysis of Model Organisms Elucidates the Association Between Environmental Factors and Human Cancer Development. International journal of molecular sciences, 25:11191, 2024
13. Takano Y, Suzuki J, Nomura K, Fujii G, Zenkoh J, Kawai H, Kuze Y, Kashima Y, Nagasawa S, Nakamura Y, Kojima M, Tsuchihara K, Seki M, Kanai A, Matsubara D, Kohno T, Noguchi M, Nakaya A, Tsuboi M, Ishii G, Suzuki Y, Suzuki A. Spatially resolved gene expression profiling of tumor microenvironment reveals key steps of lung adenocarcinoma development. Nature communications, 15:10637, 2024
14. Yamamoto G, Tanaka K, Kamata R, Saito H, Yamamori-Morita T, Nakao T, Liu J, Mori S, Yagishita S, Hamada A, Shinno Y, Yoshida T, Horinouchi H, Ohe Y, Watanabe SI, Yatabe Y, Kitai H, Konno S, Kobayashi SS, Ohashi A. WEE1 confers resistance to KRAS(G12C) inhibitors in non-small cell lung cancer. Cancer letters, 611:217414, 2024
