Annual Report 2024
Division of Cancer Immunology
Hiroyoshi Nishikawa, Kota Itahashi, Shohei Koyama, Kosuke Tanaka, Yasuko Tada, Eri Sugiyama, Yuki Koike, Kotaro Nomura, Genki Okumura, Yi-tzu Lin, Akito Fukuda, Akihito Kawazoe, Junichiro Yuda, Shugo Yajima, Chie Ozawa, Tomoka Takaku, Yoko Ohira, Shota Fukuoka, Tamiyo Kobayashi, Tomohiro Iwasawa, Kazuhiro Kumagai, Tomoya Suzuki, Yasuki Adachi, Daisuke Ito, Takuya Owari, Masaki Kondo, Fumihiro Terasaki, Naoki Yuri, Sayuri Fukaya, Takehiko Manabe, Hirotomo Machiyama, Sho Isoyama
Introduction
Cancer immunotherapy, represented by immune checkpoint inhibitors (ICIs), has become one of the standard treatment options for various types of cancer. These ICIs are frequently administered in combination with conventional drugs in clinical settings. Various biomarkers such as PD-L1 expression in tumors and tumor mutation burden (TMB) have been identified to predict therapeutic responses, demonstrating their effectiveness in specific cancers. However, our understanding of markers based on stratification by immuno-genomic analysis of the tumor microenvironment (TME) that can accurately predict the therapeutic efficacy of ICIs remains limited. In addition, as the utilization of ICI therapy expands, the number of cases of long-term response and relapse after long-term treatment is increasing, and the mechanism(s) for long-term response and the resistance to ICIs needs to be clarified.
The Team and What We Do
To address these challenges, our laboratory collaborates with clinical teams to identify novel biomarkers related to clinical responses by ICIs, to elucidate the mechanisms involved in treatment resistance to ICIs, and to propose combination treatment strategies targeting these mechanisms. We analyze tumor tissues, blood, and stools before and after various treatments including ICIs, and comprehensively analyze both cancer cells and immune cells by multicolor flow cytometry, multiplex immunohistochemistry, CyTOF, single cell RNA/ATAC/TCR seq, whole exome analysis, and metagenomic analysis (Figure 1). In addition, we investigate the mechanisms of the phenomena observed in humans by transferring them to mouse models. We are also actively involved in collaborative research with pharmaceutical companies, aiming at introducing new biomarkers into clinical trials and developing new immunotherapies and combination cancer therapies.
Figure 1. We are investigating the dynamic immune state in cancer patients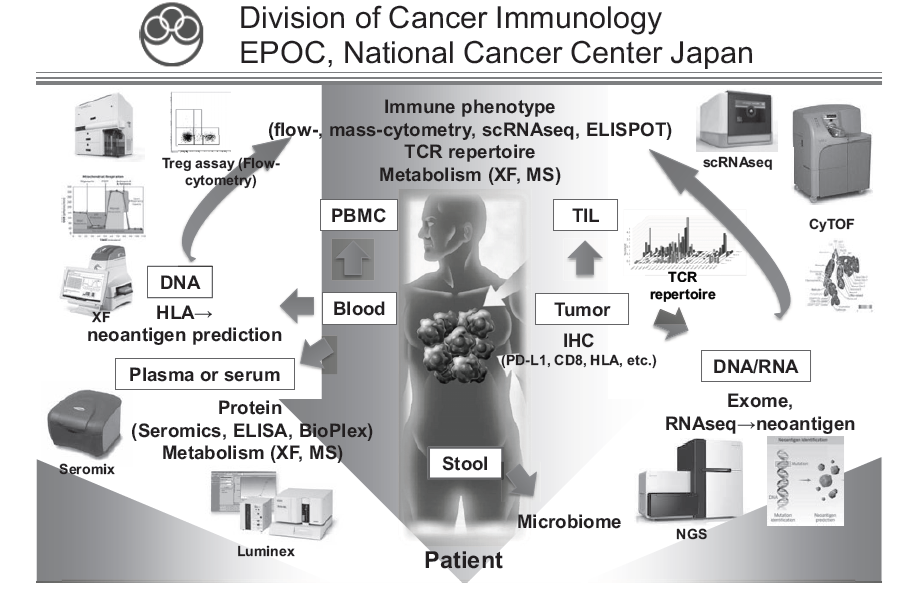

Research Activities
With the cooperation of clinical teams, tumor and stool samples from patients with non-small cell lung cancer and gastric cancer were collected to evaluate the relationship between PD-1 blockade therapy and the gut microbiota. A previously unreported bacterial strain, YB328, was successfully isolated and cultured from patients who responded to PD-1 blockade therapy. We demonstrated that YB328 activates CD103⁺ dendritic cells in the intestine. These cells then migrate to tumor tissues and draining lymph nodes to exert immune effects (Nina Yi-Tzu Lin et al. Nature 2025) (Figure 2). Furthermore, we have analyzed the mechanisms of tumor immune evasion using human specimens and cancer-bearing mouse models and have reported on the mechanisms by which driver genes and cancer signals, including EGFR gene mutations, RHOA gene mutations, MYC gene amplification, and WNT/β-catenin signaling, edit the tumor micro-immune environment in an immunosuppressive manner via cytokines and metabolites (Sugiyama Eri et al. Sci Immunol. 2020, Kumagai Shogo et al. Immunity 2020, Takeuchi Yoshiko et al. Sci Immunol. 2021, Kumagai Shogo et al. Cancer Cell 2022). Including our research, the progress made in the field of tumor immunology over the past decade has shown that some driver genes and cancer signals not only induce cancer cell survival and proliferation, but also construct immunosuppressive networks at the same time, and we have proposed the concept of “immunogenomic cancer evolution” (Kumagai Shogo et al. Nat Rev Clin Oncol. 2024; Kumagai Shogo et al. Sci Immunol. 2025). Therefore, molecularly targeted drugs that act on these driver genes and oncogenic signaling pathways may not only directly eliminate cancer cells but also disrupt the immunosuppressive tumor network, underscoring the growing importance of “immunogenomic precision medicine.”
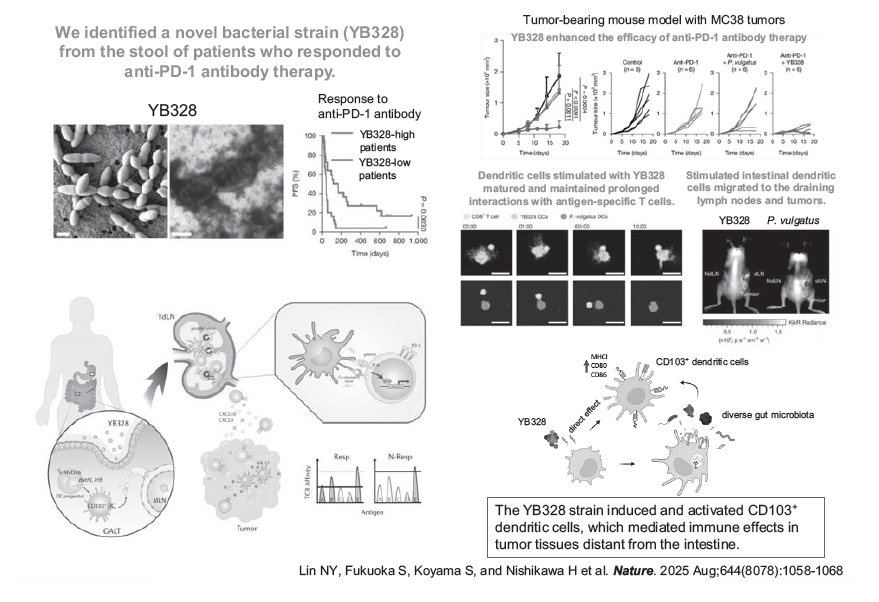
Clinical Trials
We have collaborated with several pharmaceutical companies in their clinical trials to elucidate biomarkers for predicting therapeutic responses and to evaluate immune status in the TME using our real-time immune monitoring.
Education
Many graduate school students are trained in our division, and young residents in the NCCHE are also trained to become physician scientists. After finishing their PhD course, some of the students continue their cancer immunology studies abroad.
Future Prospects
ICIs have undoubtedly opened a new era in cancer treatment. However, we still have several issues to be clarified, including acquired resistance and suppressive immune modulation by somatic mutations in cancer cells, to overcome the drawbacks of ICIs. In order to answer these questions, immunological analyses using clinical specimens are essential. Therefore, it is important to develop effective therapeutic strategies combining ICIs with other anti-cancer drugs and to identify biomarkers that can stratify responders from non-responders by clarifying the immune suppressive networks in the TME.
List of papers published in 2024
Journal
1. Tsuge A, Watanabe S, Kawazoe A, Togashi Y, Itahashi K, Masuda M, Sai A, Takei S, Muraoka H, Ohkubo S, Sugiyama D, Yan Y, Fukuoka S, Doi T, Shitara K, Koyama S, Nishikawa H. The HSP90 Inhibitor Pimitespib Targets Regulatory T Cells in the Tumor Microenvironment. Cancer immunology research, 13:273-285, 2025
2. Koganemaru S, Koyama S, Suto F, Koga M, Inaki K, Kuwahara Y, Arita T, Hirata T, Goto H, Wada N, Kobayashi M, Shibutani T, Okabayashi T, Nakamaru K, Kawazoe A, Togashi Y, Nishikawa H, Shitara K. The Tumor Immune Microenvironment and Therapeutic Efficacy of Trastuzumab Deruxtecan in Gastric Cancer. Cancer research communications, 5:84-93, 2025
3. Wagner M, Nishikawa H, Koyasu S. Reinventing type 2 immunity in cancer. Nature, 637:296-303, 2025
4. Sato T, Sugiyama D, Koseki J, Kojima Y, Hattori S, Sone K, Nishinakamura H, Ishikawa T, Ishikawa Y, Kato T, Kiyoi H, Nishikawa H. Sustained inhibition of CSF1R signaling augments antitumor immunity through inhibiting tumor-associated macrophages. JCI insight, 10:e178146, 2025
5. Moyer A, Tanaka K, Cheng EH. Apoptosis in Cancer Biology and Therapy. Annual review of pathology, 20:303-328, 2025
6. Nishinakamura H, Shinya S, Irie T, Sakihama S, Naito T, Watanabe K, Sugiyama D, Tamiya M, Yoshida T, Hase T, Yoshida T, Karube K, Koyama S, Nishikawa H. Coactivation of innate immune suppressive cells induces acquired resistance against combined TLR agonism and PD-1 blockade. Science translational medicine, 17:eadk3160, 2025
7. Bando H, Kumagai S, Kotani D, Mishima S, Irie T, Itahashi K, Tanaka Y, Habu T, Fukaya S, Kondo M, Tsushima T, Hara H, Kadowaki S, Kato K, Chin K, Yamaguchi K, Kageyama SI, Hojo H, Nakamura M, Tachibana H, Wakabayashi M, Fukui M, Fuse N, Koyama S, Mano H, Nishikawa H, Shitara K, Yoshino T, Kojima T. Atezolizumab following definitive chemoradiotherapy in patients with unresectable locally advanced esophageal squamous cell carcinoma - a multicenter phase 2 trial (EPOC1802). Nature cancer, 6:445-459, 2025
8. Kumagai S, Momoi Y, Nishikawa H. Immunogenomic cancer evolution: A framework to understand cancer immunosuppression. Science immunology, 10:eabo5570, 2025
9. Momoi Y, Kumagai S, Nishikawa H. Immunogenomic precision medicine: a personalized approach based on immunogenomic cancer evolution. International immunology, dxaf020, 2025
10. Jinushi K, Saito T, Kurose K, Suzuki S, Kojima T, Takahara T, Makino T, Ogawa T, Nishikawa H, Kakimi K, Iida S, Nakajima J, Doki Y, Oka M, Ueda R, Wada H. Phase I study on neoadjuvant combination immunotherapy with mogamulizumab and nivolumab for solid tumors. Journal for immunotherapy of cancer, 13:e010634, 2025
11. Terasaki F, Sugiura T, Okamura Y, Ashida R, Ohgi K, Yamada M, Ohtsuka S, Uesaka K. Benefit of lymph node dissection for perihilar and distal cholangiocarcinoma according to lymph node stations. Journal of hepato-biliary-pancreatic sciences, 31:251-261, 2024
12. Nishikawa H. Establishment of immune suppression by cancer cells in the tumor microenvironment. Proceedings of the Japan Academy. Series B, Physical and biological sciences, 100:114-122, 2024
13. Kumagai S, Itahashi K, Nishikawa H. Regulatory T cell-mediated immunosuppression orchestrated by cancer: towards an immuno-genomic paradigm for precision medicine. Nature reviews. Clinical oncology, 21:337-353, 2024
14. Fukuda A, Okuma Y. From Rarity to Reality: Osimertinib's Promising Horizon in Treating Uncommon EGFR Mutations in Non-Small Cell Lung Cancer. Clinical cancer research, 30:3128-3136, 2024
15. Yamamoto G, Tanaka K, Kamata R, Saito H, Yamamori-Morita T, Nakao T, Liu J, Mori S, Yagishita S, Hamada A, Shinno Y, Yoshida T, Horinouchi H, Ohe Y, Watanabe SI, Yatabe Y, Kitai H, Konno S, Kobayashi SS, Ohashi A. WEE1 confers resistance to KRAS(G12C) inhibitors in non-small cell lung cancer. Cancer letters, 611:217414, 2024
