Home > Division of Cancer Therapeutics
Division of Cancer Therapeutics
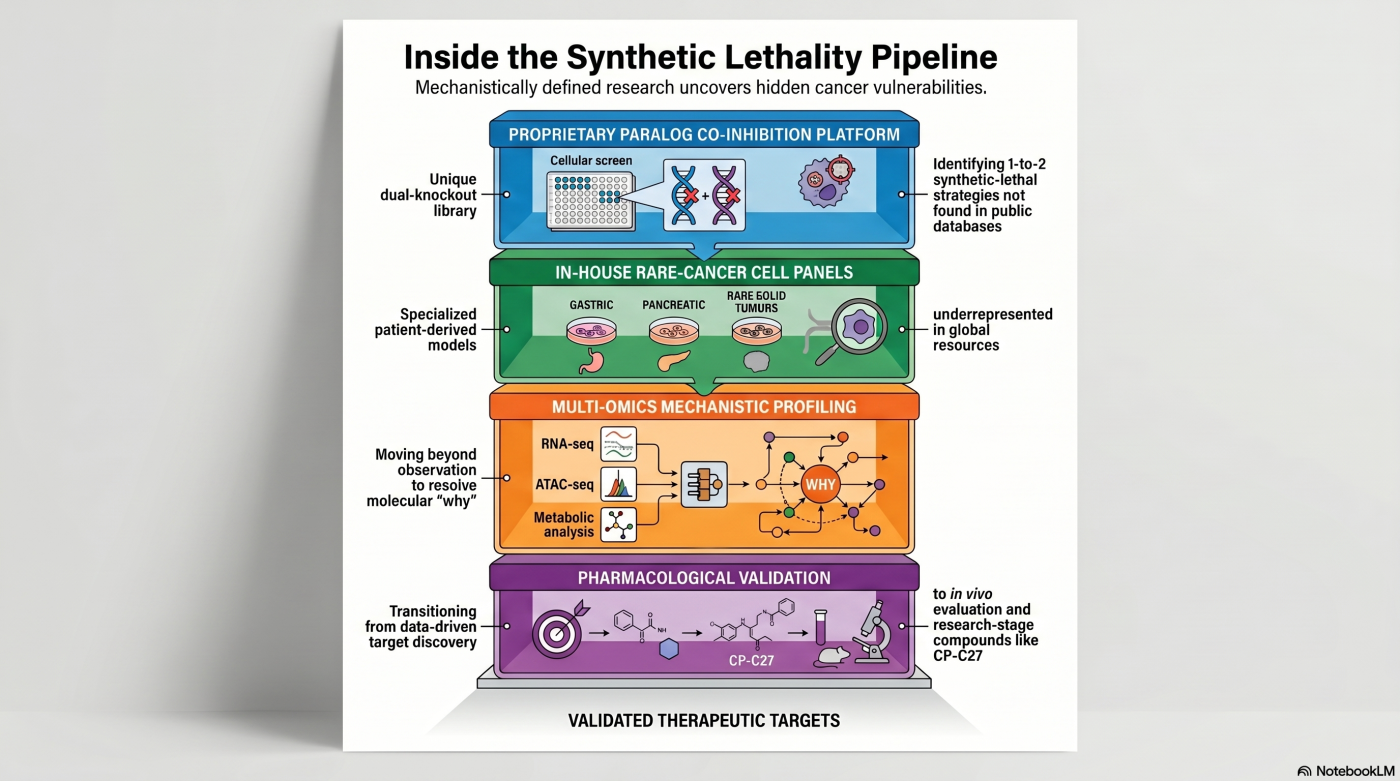
Original Concepts. Mechanistically Defined Cancer Vulnerabilities.
Division of Cancer Therapeutics, National Cancer Center Research Institute
The Division of Cancer Therapeutics pursues synthetic-lethality-based target discovery in genetically defined refractory cancers — including SWI/SNF-deficient, KRAS-mutant, and SMAD4 / KDM6A-deficient tumors. We integrate target identification, mechanistic dissection, compound evaluation, and early drug-discovery validation within a single end-to-end pipeline.
Our approach combines a proprietary paralog co-inhibition platform, in-house rare- and refractory-cancer cell-line panels, data-driven dependency analysis, and multi-omics mechanistic profiling — surfacing therapeutic targets that public-database analysis alone cannot reach.
Our Vision
Unraveling Synthetic Lethality at the Mechanistic Level.
We move beyond the observation that something "works" to resolve why it works at the molecular level. Mechanistic resolution, in our view, is the foundation of reproducible drug discovery — the point at which a vulnerability becomes a validated therapeutic target.
Our Research Foundations
Four platforms anchor our discovery pipeline, from target identification through mechanism to pharmacological evaluation.
- Paralog Co-Inhibition Platform — 1-to-2 synthetic-lethal strategy and proprietary paralog-pair dual-knockout library
- Rare- and Refractory-Cancer Cell-Line Panels — in-house disease models for SWI/SNF-deficient, KRAS-mutant, and SMAD4 / KDM6A-deficient cancers
- Data-Driven Target Discovery — DepMap dependency reanalysis integrated with in-house CRISPR/Cas9 screening
- Mechanistic and Pharmacological Validation — multi-omics profiling (RNA-seq, ATAC-seq, CUT&RUN), metabolic analysis, and in vitro / in vivo pharmacological evaluation
→ See Research Projects for the full platform architecture and disease-focused programs.
Disease Focus
We target cancers in which a clear genetic vulnerability is present but molecularly targeted treatment options remain limited.
|
Category |
Representative Cancers |
Key Molecular Alterations |
|---|---|---|
|
Rare, pediatric, and AYA cancers |
Malignant rhabdoid tumor, epithelioid sarcoma, synovial sarcoma |
SMARCB1 loss, SS18-SSX fusion |
|
Refractory gynecologic and gastrointestinal cancers |
Ovarian clear cell carcinoma, diffuse-type gastric cancer (including scirrhous gastric cancer) |
ARID1A loss, SMAD4 loss |
|
Refractory solid tumors |
Non-small cell lung cancer, pancreatic cancer, esophageal cancer, renal cell carcinoma |
SMARCA4 loss, PBRM1 loss, KDM6A loss, SMAD4 loss, KRAS mutations |
Across these diseases, a defined molecular alteration coexists with limited targeted-therapy options. We identify those vulnerabilities and validate them as therapeutic targets through our discovery platforms.
→ See Research Projects for disease-focused programs.
Research Highlights
Our research is organized into six thematic areas spanning the laboratory's intellectual trajectory from conventional synthetic lethality to higher-order frameworks. Detailed deep dives for all six themes are consolidated on the Research Highlights page.
1. Conventional Synthetic Lethality
1-to-1 synthetic-lethality between paralog pairs — SMARCA2 dependency in SMARCA4-deficient cancers, and EP300 dependency in CBP-deficient cancers.
2. Paralog Co-Inhibition
1-to-2 synthetic-lethality strategy — simultaneous CBP/p300 inhibition in SMARCB1-deficient and cBAF-deficient cancers.
3. Next-Generation Synthetic Lethality
2-to-1 and 2-to-n synthetic-lethality frameworks (higher-order synthetic lethality) — for example, CHD3 dependency in dual SMARCA4 / SMARCA2-deficient cancers.
4. Data-Driven Target Discovery
Integration of DepMap dependency reanalysis with our in-house cell-line panels — including USP8 / FGFR2 dependency in ARID1A-deficient ovarian clear cell carcinoma.
5. Glutathione Metabolic Vulnerability
Glutathione metabolic dependency in SWI/SNF-deficient cancers (ARID1A, SMARCB1) — GCLC inhibition as a ferroptosis-inducing therapeutic strategy.
6. Drug Repositioning
Biomarker-defined repositioning of existing drugs — gemcitabine in ARID1A-deficient ovarian clear cell carcinoma and diffuse-type gastric cancer, exploiting a pyrimidine metabolic vulnerability.
Track Record
Our accumulated work is summarized along three axes: peer-reviewed research output, drug-discovery seeds, and industry partnerships.
Selected Publications
The three studies below define the laboratory's intellectual trajectory. The complete publication list is on the Publications page.
- Sasaki M, Kato D, Murakami K, Yoshida H, Takase S, Otsubo T, Ogiwara H*. Targeting dependency on a paralog pair of CBP/p300 against de-repression of KREMEN2 in SMARCB1-deficient cancers. Nat Commun. 2024;15(1):4770. Representative study establishing paralog co-inhibition as a synthetic-lethal strategy in SMARCB1-deficient cancers. → PubMed
- Ogiwara H*, Takahashi K, Sasaki M, Kuroda T, Yoshida H, Watanabe R, Maruyama A, Makinoshima H, Chiwaki F, Sasaki H, Kato T, Okamoto A, Kohno T*. Targeting the Vulnerability of Glutathione Metabolism in ARID1A-Deficient Cancers. Cancer Cell. 2019;35(2):177–190.e8. Demonstrated glutathione metabolic dependency in ARID1A-deficient cancers — foundation of the metabolic-vulnerability program. → PubMed
- Ogiwara H, Sasaki M, Mitachi T, Oike T, Higuchi S, Tominaga Y, Kohno T*. Targeting p300 addiction in CBP-deficient cancers causes synthetic lethality via apoptotic cell death due to abrogation of MYC expression. Cancer Discov. 2016;6(4):430–445. Established the CREBBP/EP300 paralog-dependency axis — conceptual origin of the laboratory's synthetic-lethality program. → PubMed
Published Research Compounds
Our peer-reviewed publications include the following research-stage compounds:
- CP-C27 — dual CBP/p300 inhibitor (Sasaki et al., Nat Commun 2024)
- GCLCi1 / GCLCi0 — GCLC inhibitors developed and evaluated in collaboration with Ono Pharmaceutical Co., Ltd. (Takeuchi et al., Cancer Res 2026)
These are research-use compounds disclosed in the corresponding publications.
Industry Partnerships and Public Outreach
We engage with industry partners through publication-disclosed collaborations — for example, the GCLC inhibitor program with Ono Pharmaceutical Co., Ltd. (Takeuchi et al., Cancer Res 2026). Key findings have been communicated through NCC press releases at major publication milestones.
→ See Publications for the Selected Publications Overview organized by the six research themes.
Selected Press Releases
- March 2026 — Cancer Res paper: GCLC inhibitors induce ferroptosis in SMARCB1-deficient rare cancers via a redox-homeostasis vulnerability. → NCC press release (Japanese)
- June 2024 — Nat Commun paper: A paralog co-inhibition strategy targeting CBP/p300 in SMARCB1-deficient pediatric and AYA cancers. → NCC press release (Japanese)
- January 2019 — Cancer Cell paper: Glutathione metabolism as a therapeutic vulnerability of ARID1A-deficient cancers. → NCC press release (Japanese)
- December 2015 — Cancer Discov paper: A synthetic-lethality-based therapeutic strategy for CBP-mutant cancers, including small-cell lung cancer and malignant lymphoma. → NCC press release (Japanese)
For Industry and Academic Partners
Our partnership offering is anchored by:
- A proprietary paralog-pair dual-knockout library — not represented in public dependency databases.
- In-house rare- and refractory-cancer cell-line panels — including patient-derived cell lines for gastric and pancreatic cancers, and comprehensive panels spanning lung cancer, sarcoma, esophageal cancer, and renal cancer; covering cancer types and molecular subtypes underrepresented in international public databases.
- End-to-end translational evaluation — from target identification, through mechanistic dissection, to pharmacological validation and indication-expansion analysis.
→ Contact for collaboration inquiries
Site Navigation
For additional inquiries, including media requests, please see Contact and Access.
Last Updated: 2026-05-19

